|
12/12/2023 0 Comments Apply compensation matrix flowjo 
figure ( figsize = ( 8, 4 )) ax = pylab. rename_node ( 't1', 'compensated' ) fig = pylab. rename_node ( 't1', 'uncompensated' ) data. logicle () # logicle the data so it looks more like you are used to seeing data. loadFCS ( 'E6901F0T-07_CMV pp65.fcs', auto_comp = False, transform = None ) data. load_compensate_matrix ( 'CompMatrixDenny06Nov09' ) data = fcm. Import fcm import fcm.graphics as graph import matplotlib.pyplot as pylab sidx, comp = fcm. Of working with the FCMdata data tree which will be covered in the next section Logicle transforms and compensation to a FCMdata object. The code below shows how to control and manually apply The transform argument of log or automatic transformation can be prevented by setting theįurther FCMdata provides pensate(), FCMdata.logicle(),Īnd FCMdata.log() methods. The log transform can be used instead by passing By default when loading anįcs file loadFCS() will apply the logicle transform to all fluorescent channels withĪ range of 262144 (PNR in the fcs header). Return the laser names (sidx) and compensation matrix exported in the format used by Flowjo.įcm also supports the logicle and log data transforms. You to provide your own compensation matrix by passing in the comp and sidx arguments, orĬan not compensate by passing False as the auto_comp argument to loadFCS().įor convenience fcm provides the load_compensate_matrix() which will By default loadFCS() compensates fcsĭata using the compensation matrix provided in fcs file header. To try and counter fluorescent spill over. The matrix will be applied to the group assuming all parameters are the same.Along with reading FCS files, fcm provides methods to apply compensation This matrix file can also be dragged to the group. Notice the sample A1 above now has a different color badge from the matrix we dragged on it. xml matrix file from your file system onto a sample in FJ to compensate that sample: You can apply a matrix stored on your file system to samples in your workspace assuming they have all the parameters listed in the matrix itself. The names must match exactly best practice is to use the names that appear in the $PnN keywords of the FCS file. Parameters/channels that are not in this list have not been compensated parameters/channels in this list that are not in the sample are ignored. The file contains the title of the compensation matrix, the names of the parameters/channels used in the matrix and the coefficients of the compensated parameters. The saved matrix file is written in standard XML and is based on standards developed by the Data Standards Task Force (DSTF) of the International Society for the Advancement of Cytometry (ISAC). 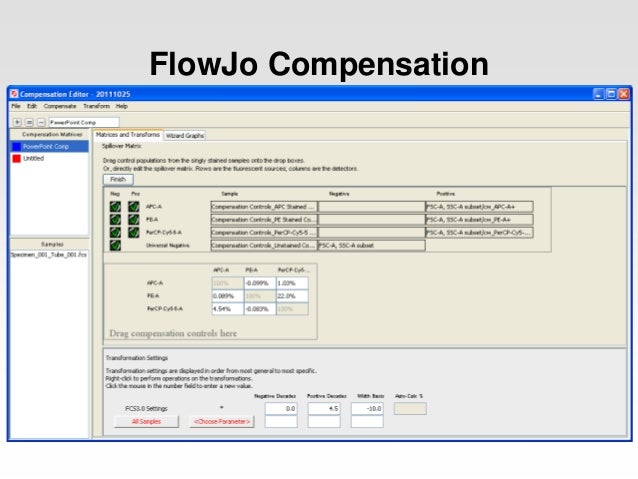
FlowJo uses XML to express these matrices.Įxample matrix file, viewed in Mozilla Firefox browser. Once in the Matrix Editor, you can use the File -> Save Matrix command to export it. 
The matrix editor is easily accessed by double-clicking a sample’s compensation badge (the icon looks like a tic-tac-toe grid): 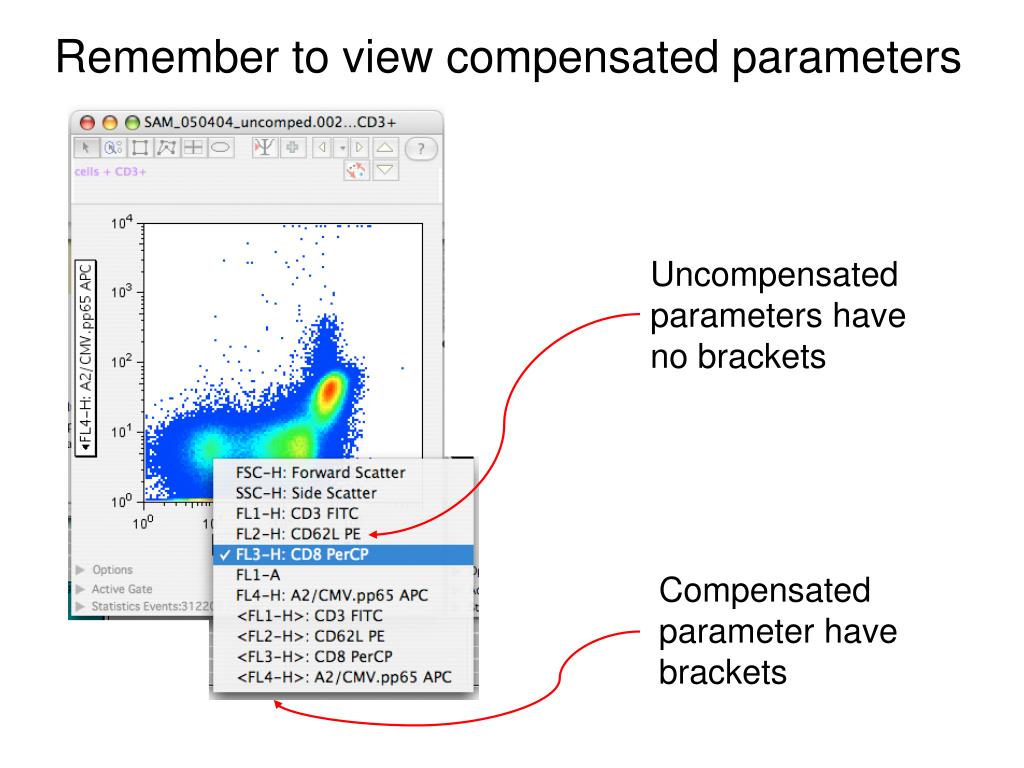
To save a matrix to its own stand-alone file, use the Matrix Editor. Whether compensation comes from your data as Acquisition-Defined, or is generated via FlowJo’s compensation wizard, the resulting matrix of values will be stored in XML in the FlowJo workspace document. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
